![]()
Do sports problems require tailored methods or direct applications?
On a swimming-oriented journey with a Bayesian roadmap.
Arthur Leroy - Department of Computer Science, The University of Manchester
S-Training seminar - 24/02/2023
First steps in the world of sports sciences
Which of the 5 disciplines of modern pentathlon has the most impact on performance?
When looking directly at raw data from international competitions, it was clear that swimming carried the most points in total, although swimming specialists often felt disadvantaged.

Modern pentathlon: additional insights


\(\rightarrow\) You sometimes need to change the question to identify the correct answer from data.
Choosing your battles is key
In general, it is crucial to identify which problem and tools we have at hand:
- Type 1: Classical methods work just fine on our data.
- Type 2: Classical methods are not adapted to the context, our data, etc.
- Type 3: The information needed to solve our problem is not contained is our data.

How can we make life of our colleagues easier?
While tackling type 1 problems properly allows statisticians to quickly conduct fruitful collaborations, it is probably on type 2 that they can achieve the most valuable contributions.
\(\rightarrow\) Since it is legitimately frustrating to encounter type 2 problems for any applied scientist, it can be tempting to tweak existing methods or force a square peg into a round hole.

Consequences of frustration: simplicity is sometimes limited
\[ACWR(N) = \frac{L^{week}_N}{L^{week}_N + L^{week}_{N-7} + L^{week}_{N-14} + L^{week}_{N-21}}\]
\[EWMA(N) = \frac{2}{N+1} L(N) + (1 - \frac{2}{N+1}) EWMA(N-1)\]

Make life easier: simplicity might also work great
\[REDI_{\lambda}(N) = \frac{1}{\sum\limits_{t = 1}^{N} e^{- \lambda t}} \sum\limits_{t = 1}^{N} e^{- \lambda t} L(t)\]

Make life easier: simplicity might also work great
Moussa, Leroy et al. - Robust Exponential Decreasing Index (REDI): adaptive and robust method for computing cumulated workload - BMJ SEM - 2019

Consequences of frustration: good intentions, bad ideas
Hopkins - Estimating Sample Size for Magnitude-Based Inferences - Sport science - 2006
As for any discipline involving data collection on humans, sports sciences often had to deal with low sample sizes, resulting in difficulties to identify meaningful effects using traditional statistical tools (while also raising legitimate concerns about standard NHST). In this context, some authors with more interest in statistics tried to provide DIY Excel spreadsheets that somehow became the field’s standard.

Consequences of frustration: controversial scientific practices
Sainani - The Problem with “Magnitude-based Inference” - Med Sci Sports Exerc - 2018
In a series of articles and presentations, Kristin Sainani (and several others) identified and highlighted serious concerns about MBI and its use (exclusively) in sports sciences. The responses from authors and influential people have been particularly virulent and exhibited major issues in the field, from honest and naive misuse to apparent scientific misconducts.

Life is easy(-ish): the wheel has already been invented


Type 1 case study: influence of morphology in swimming
Pla, Leroy et al. - Bayesian approach to quantify morphological impact on performance in international elite freestyle swimming - BMJ SEM - 2019
To study the influence of morphology on the performance in swimming, we can simply leverage a Bayesian hierarchical mixed-model like:
\[S_i = c_0 + c_H \times H + c_W \times W + \gamma_i + \varepsilon,\]
where we assume appropriate priors on \(\{ c_0, c_H, c_W, \gamma_i \}\), and
- \(S_i\) is the speed of the swimmer in \(m/s\),
- \(H\) is the height of the swimmer,
- \(W\) is the weight of the swimmer.
Type 1 case study: influence of morphology in swimming


Type 1 case study: improvement over successive events
Pla, Leroy et al. - International Swimming League: Do Successive Events Lead to Improve Swimming Performance? - IJSPP - 2021


Some problems are genuinely hard: performance of swimmers

- Irregular time series (in number of observations and location),
- Many different swimmers per category,
- A few observations per swimmer.
Application of existing methods: promising but limited
Leroy et al. - Functional Data Analysis in Sport Science: Example of Swimmers’ Progression Curves Clustering - Applied Sciences - 2018
Functional data analysis is a field that model continuous phenomenons as random functions. By using B-splines and a simple k-means algorithm, we can identify clusters of functions:

\(\rightarrow\) However, this comes with serious limitations in terms modelling and prediction.
Moving beyond direct application: Gaussian process regression
\[y = \color{orange}{f}(x) + \epsilon\]
No restrictions on \(\color{orange}{f}\) but a prior distribution on a functional space: \(\color{orange}{f} \sim \mathcal{GP}(0,C(\cdot,\cdot))\)

- Powerful non parametric method offering probabilistic predictions,
- Computational complexity in \(\mathcal{O}(N^3)\), with N the number of observations.
Modelling and prediction with a unique GP

GPs are great for modelling time series although insufficient for long-term predictions.
Multi-task GP with common mean (Magma)
Leroy et al. - MAGMA: Inference and Prediction using Multi-Task Gaussian Processes with Common Mean - Machine Learning - 2022
\[y_i = \mu_0 + f_i + \epsilon_i\]
with:
- \(\mu_0 \sim \mathcal{GP}(m_0, K_{\theta_0}),\)
- \(f_i \sim \mathcal{GP}(0, \Sigma_{\theta_i}), \ \perp \!\!\! \perp_i,\)
- \(\epsilon_i \sim \mathcal{GP}(0, \sigma_i^2), \ \perp \!\!\! \perp_i.\)
It follows that:
\[y_i \mid \mu_0 \sim \mathcal{GP}(\mu_0, \Sigma_{\theta_i} + \sigma_i^2 I), \ \perp \!\!\! \perp_i\]
\(\rightarrow\) Unified GP framework with a common mean process \(\mu_0\), and individual-specific process \(f_i\),
\(\rightarrow\) Naturaly handles irregular grids of input data.
EM algorithm
E step: \[ \begin{align} p(\mu_0(\mathbf{t}) \mid \textbf{y}, \hat{\Theta}) &\propto \mathcal{N}(\mu_0(\mathbf{t}); m_0(\textbf{t}), \textbf{K}_{\hat{\theta}_0}^{\textbf{t}}) \times \prod\limits_{i =1}^M \mathcal{N}(\mathbf{y}_i; \mu_0( \textbf{t}_i), \boldsymbol{\Psi}_{\hat{\theta}_i, \hat{\sigma}_i^2}^{\textbf{t}_i}) \\ &= \mathcal{N}(\mu_0(\mathbf{t}); \hat{m}_0(\textbf{t}), \hat{\textbf{K}}^{\textbf{t}}), \end{align} \] M step: \[ \begin{align*} \hat{\Theta} &= \underset{\Theta}{\arg\max} \ \ \log \mathcal{N} \left( \hat{m}_0(\textbf{t}); m_0(\textbf{t}), \mathbf{K}_{\theta_0}^{\textbf{t}} \right) - \dfrac{1}{2} Tr \left( \hat{\mathbf{K}}^{\textbf{t}} {\mathbf{K}_{\theta_0}^{\textbf{t}}}^{-1} \right) \\ & \ \ \ + \sum\limits_{i = 1}^{M}\left\{ \log \mathcal{N} \left( \mathbf{y}_i; \hat{m}_0(\mathbf{t}_i), \boldsymbol{\Psi}_{\theta_i, \sigma^2}^{\mathbf{t}_i} \right) - \dfrac{1}{2} Tr \left( \hat{\mathbf{K}}^{\mathbf{t}_i} {\boldsymbol{\Psi}_{\theta_i, \sigma^2}^{\mathbf{t}_i}}^{-1} \right) \right\}. \end{align*} \]
Prediction
For a new individual, we observe
some data \(y_*(\textbf{t}_*)\). Let us
recall:
\[y_* \mid \mu_0 \sim \mathcal{GP}(\mu_0, \boldsymbol{\Psi}_{\theta_*, \sigma_*^2}), \ \perp \!\!\! \perp_i\]
Goals:
- derive a analytical predictive distribution at arbitrary inputs \(\mathbf{t}^{p}\),
- sharing the information from training individuals, stored in the mean process \(\mu_0\).
Difficulties:
- the model is conditionned over \(\mu_0\), a latent, unobserved quantity,
- defining the adequate target distribution is not straightforward,
- working on a new grid of inputs \(\mathbf{t}^{p}_{*}= (\mathbf{t}_{*}, \mathbf{t}^{p})^{\intercal},\) potentially distinct from \(\mathbf{t}.\)
Prediction: the key idea
Defining a multi-task prior distribution by:
- conditioning on training data,
- integrating over \(\mu_0\)’s hyper-posterior distribution.
\[\begin{align} p(y_* (\textbf{t}_*^{p}) \mid \textbf{y}) &= \int p\left(y_* (\textbf{t}_*^{p}) \mid \textbf{y}, \mu_0(\textbf{t}_*^{p})\right) p(\mu_0 (\textbf{t}_*^{p}) \mid \textbf{y}) \ d \mu_0(\mathbf{t}^{p}_{*}) \\ &= \int \underbrace{ p \left(y_* (\textbf{t}_*^{p}) \mid \mu_0 (\textbf{t}_*^{p}) \right)}_{\mathcal{N}(y_*; \mu_0, \Psi_*)} \ \underbrace{p(\mu_0 (\textbf{t}_*^{p}) \mid \textbf{y})}_{\mathcal{N}(\mu_0; \hat{m}_0, \hat{K})} \ d \mu_0(\mathbf{t}^{p}_{*}) \\ &= \mathcal{N}( \hat{m}_0 (\mathbf{t}^{p}_{*}), \Gamma) \end{align}\]
Prediction: additional steps
-
Multi-task prior:
\[p \left( \begin{bmatrix} y_*(\color{grey}{\mathbf{t}_{*}}) \\ y_*(\color{purple}{\mathbf{t}^{p}}) \\ \end{bmatrix} \mid \textbf{y} \right) = \mathcal{N} \left( \begin{bmatrix} y_*(\color{grey}{\mathbf{t}_{*}}) \\ y_*(\color{purple}{\mathbf{t}^{p}}) \\ \end{bmatrix}; \ \begin{bmatrix} \hat{m}_0(\color{grey}{\mathbf{t}_{*}}) \\ \hat{m}_0(\color{purple}{\mathbf{t}^{p}}) \\ \end{bmatrix}, \begin{pmatrix} \Gamma_{\color{grey}{**}} & \Gamma_{\color{grey}{*}\color{purple}{p}} \\ \Gamma_{\color{purple}{p}\color{grey}{*}} & \Gamma_{\color{purple}{pp}} \end{pmatrix} \right)\]
-
Multi-task posterior:
\[p(y_*(\color{purple}{\mathbf{t}^{p}}) \mid y_*(\color{grey}{\mathbf{t}_{*}}), \textbf{y}) = \mathcal{N} \Big( y_*(\color{purple}{\mathbf{t}^{p}}); \ \hat{\mu}_{*}(\color{purple}{\mathbf{t}^{p}}) , \hat{\Gamma}_{\color{purple}{pp}} \Big)\]
with:
- \(\hat{\mu}_{*}(\color{purple}{\mathbf{t}^{p}}) = \hat{m}_0(\color{purple}{\mathbf{t}^{p}}) + \Gamma_{\color{purple}{p}\color{grey}{*}}\Gamma_{\color{grey}{**}}^{-1} (y_*(\color{grey}{\mathbf{t}_{*}}) - \hat{m}_0 (\color{grey}{\mathbf{t}_{*}}))\)
- \(\hat{\Gamma}_{\color{purple}{pp}} = \Gamma_{\color{purple}{pp}} - \Gamma_{\color{purple}{p}\color{grey}{*}}\Gamma_{\color{grey}{**}}^{-1} \Gamma_{\color{grey}{*}\color{purple}{p}}\)
This multi-task posterior distribution provides the desired probabilistic predictions.
A GIF is worth a thousand words
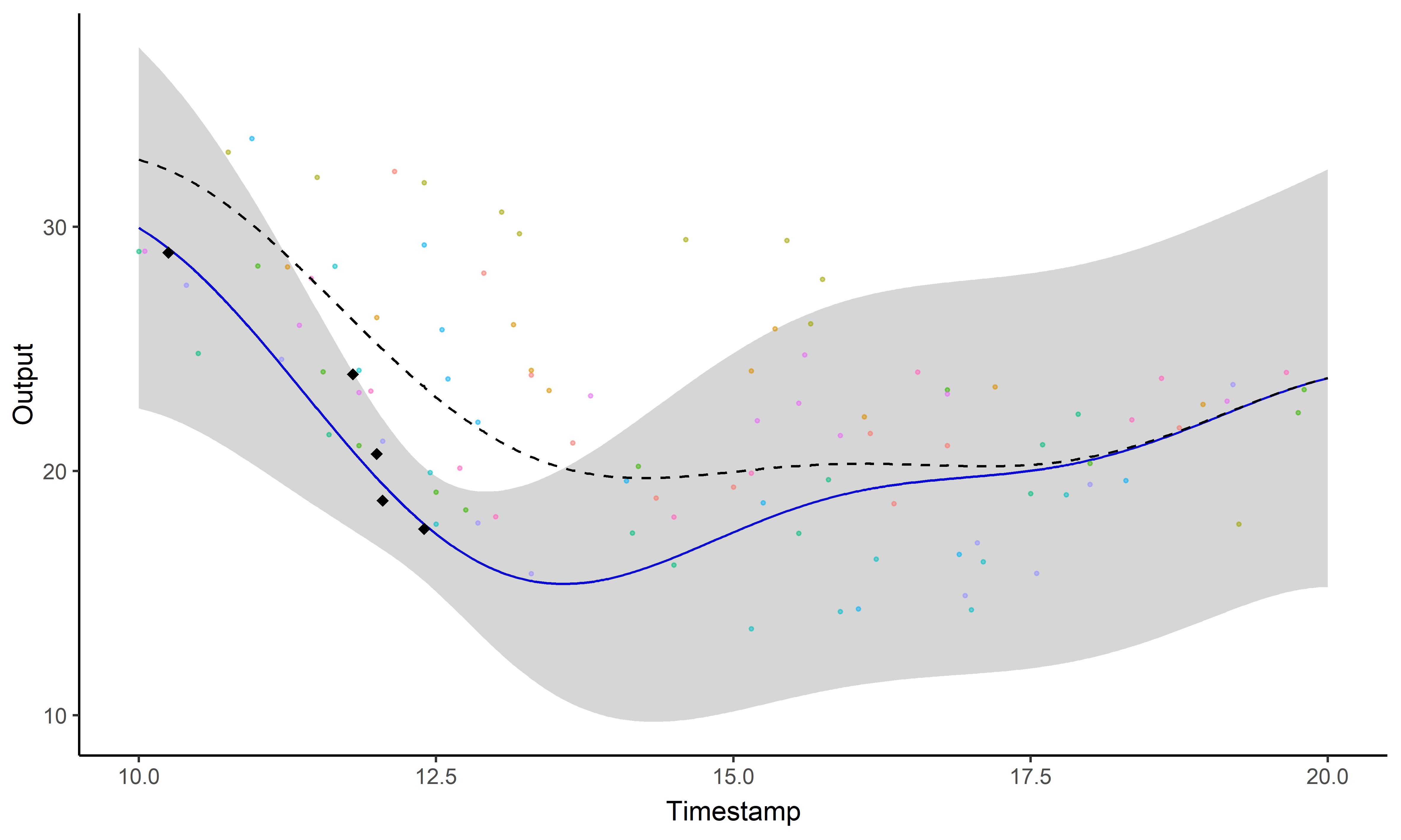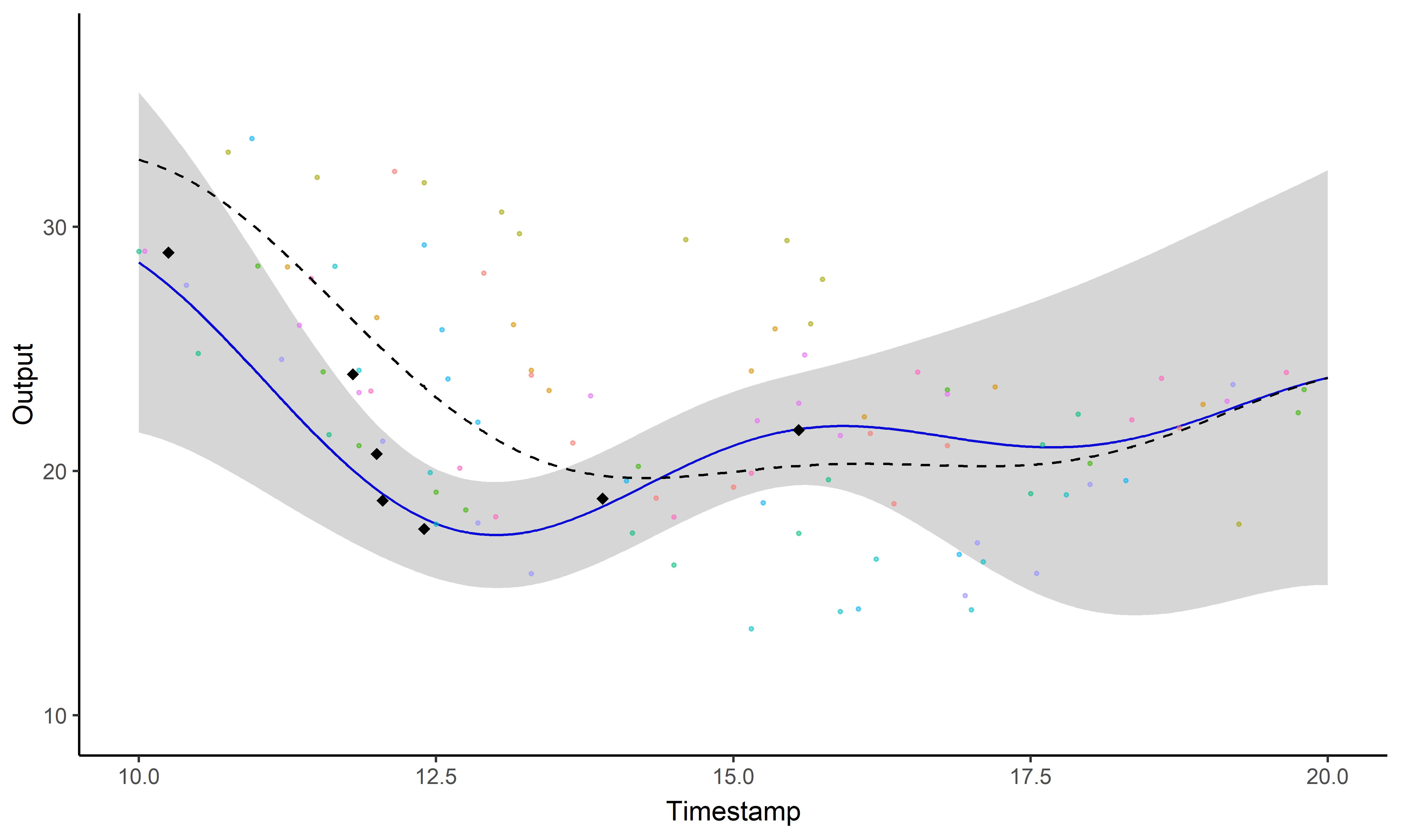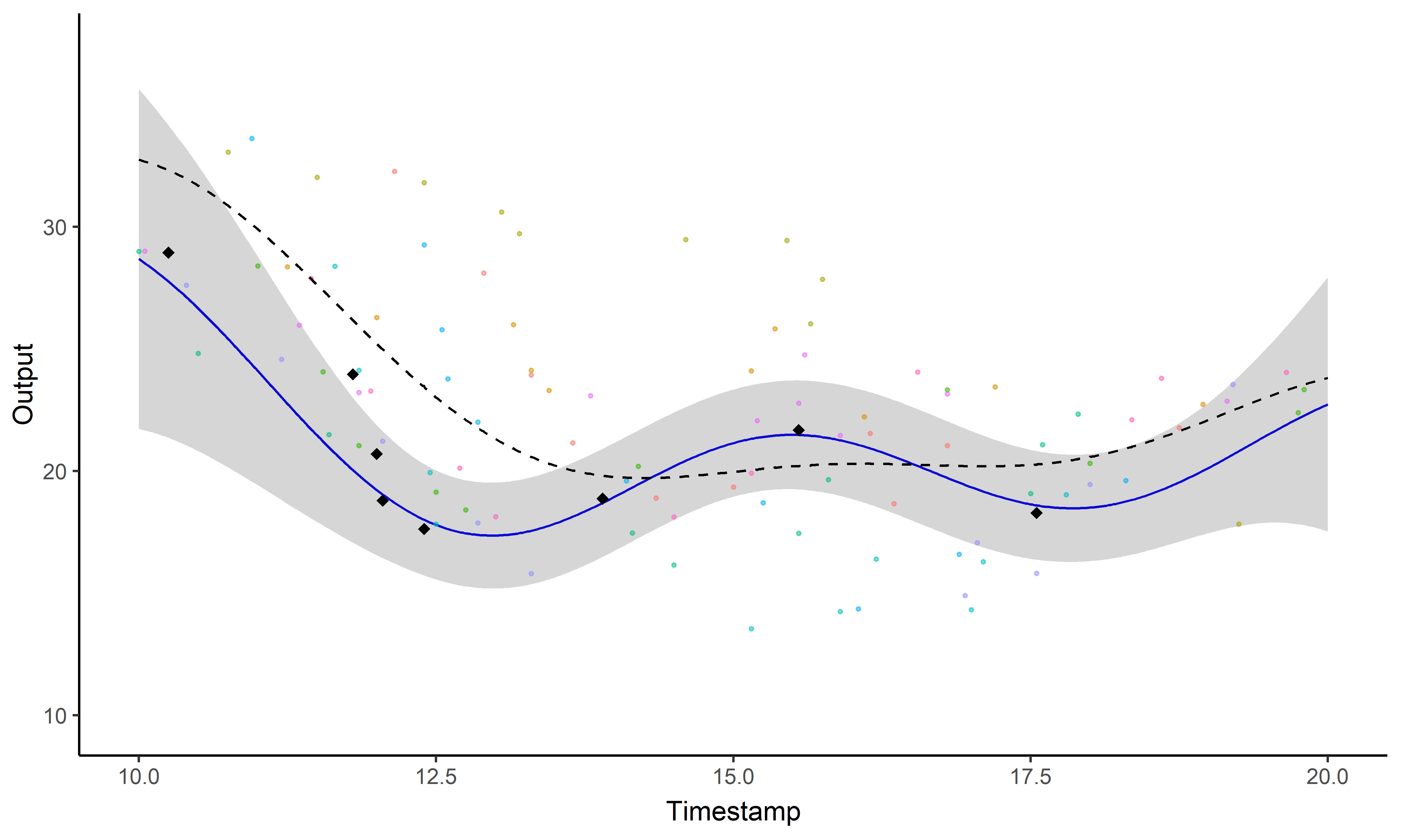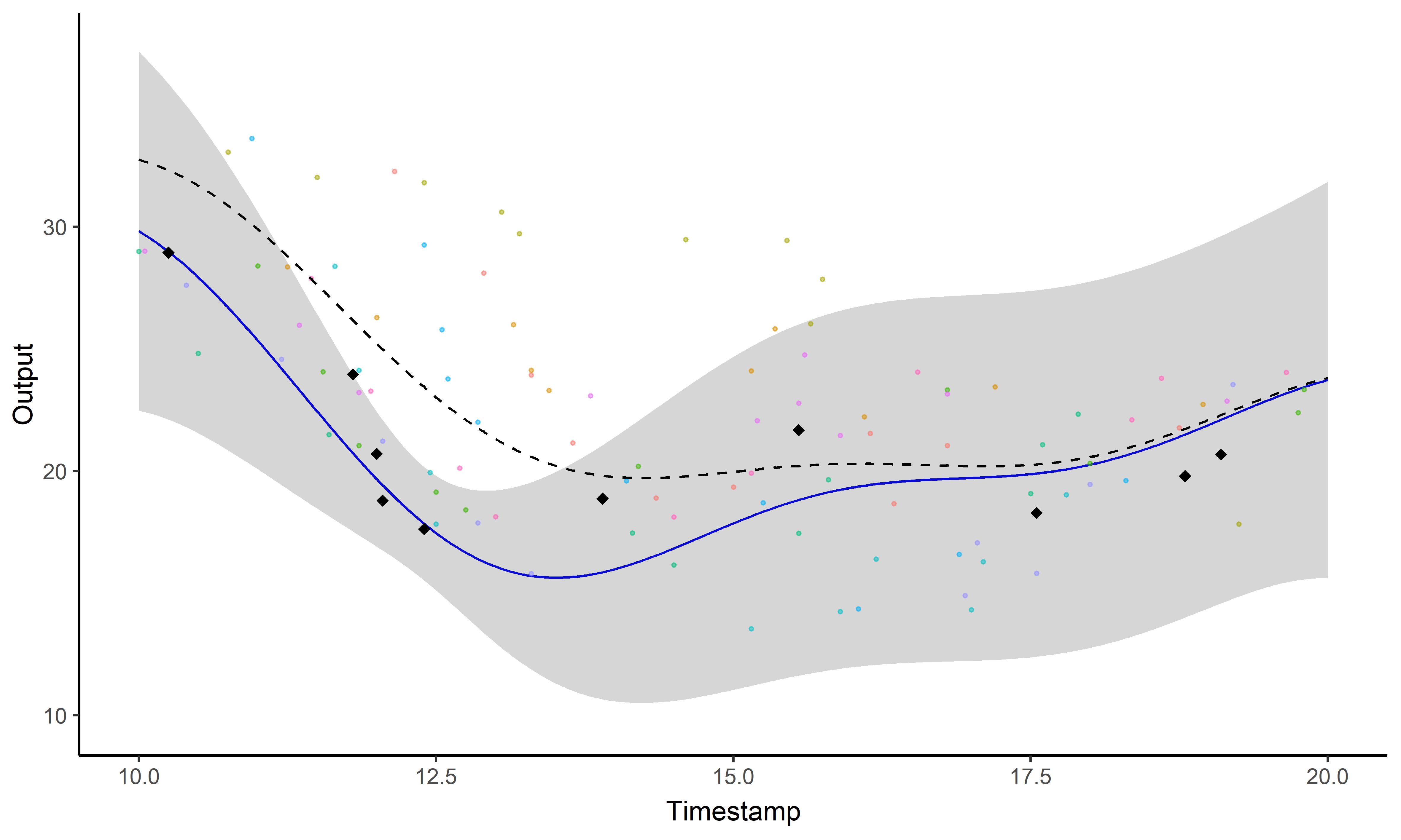
Magma + Clustering = MagmaClust
Leroy et al. - Cluster-Specific Predictions with Multi-Task Gaussian Processes - Journal of Machine Learning Research - 2023
A unique underlying mean process might be too restrictive.
\(\rightarrow\) Mixture of multi-task GPs:
\[y_i = \mu_0 + f_i + \epsilon_i\]
with:
- \(\color{green}{Z_{i}} \sim \mathcal{M}(1, \color{green}{\boldsymbol{\pi}}), \ \perp \!\!\! \perp_i,\)
- \(\mu_0 \sim \mathcal{GP}(m_0, K_{\theta_0}), \ \perp \!\!\! \perp_k,\)
- \(f_i \sim \mathcal{GP}(0, \Sigma_{\theta_i}), \ \perp \!\!\! \perp_i,\)
- \(\epsilon_i \sim \mathcal{GP}(0, \sigma_i^2), \ \perp \!\!\! \perp_i.\)
It follows that:
\[y_i \mid \mu_0 \sim \mathcal{GP}(\mu_0, \Psi_i), \ \perp \!\!\! \perp_i\]
Magma + Clustering = MagmaClust
Leroy et al. - Cluster-Specific Predictions with Multi-Task Gaussian Processes - Journal of Machine Learning Research - 2023
A unique underlying mean process might be too restrictive.
\(\rightarrow\) Mixture of multi-task GPs:
\[y_i \mid \{\color{green}{Z_{ik}} = 1 \} = \mu_{\color{green}{k}} + f_i + \epsilon_i\]
with:
- \(\color{green}{Z_{i}} \sim \mathcal{M}(1, \color{green}{\boldsymbol{\pi}}), \ \perp \!\!\! \perp_i,\)
- \(\mu_{\color{green}{k}} \sim \mathcal{GP}(m_{\color{green}{k}}, \color{green}{C_{\gamma_{k}}})\ \perp \!\!\! \perp_{\color{green}{k}},\)
- \(f_i \sim \mathcal{GP}(0, \Sigma_{\theta_i}), \ \perp \!\!\! \perp_i,\)
- \(\epsilon_i \sim \mathcal{GP}(0, \sigma_i^2), \ \perp \!\!\! \perp_i.\)
It follows that:
\[y_i \mid \mu_0 \sim \mathcal{GP}(\mu_0, \Psi_i), \ \perp \!\!\! \perp_i\]
Magma + Clustering = MagmaClust
A unique underlying mean process might be too restrictive.
\(\rightarrow\) Mixture of multi-task GPs:
\[y_i \mid \{\color{green}{Z_{ik}} = 1 \} = \mu_{\color{green}{k}} + f_i + \epsilon_i\]
with:
- \(\color{green}{Z_{i}} \sim \mathcal{M}(1, \color{green}{\boldsymbol{\pi}}), \ \perp \!\!\! \perp_i,\)
- \(\mu_{\color{green}{k}} \sim \mathcal{GP}(m_{\color{green}{k}}, \color{green}{C_{\gamma_{k}}})\ \perp \!\!\! \perp_{\color{green}{k}},\)
- \(f_i \sim \mathcal{GP}(0, \Sigma_{\theta_i}), \ \perp \!\!\! \perp_i,\)
- \(\epsilon_i \sim \mathcal{GP}(0, \sigma_i^2), \ \perp \!\!\! \perp_i.\)
It follows that:
\[y_i \mid \{ \boldsymbol{\mu} , \color{green}{\boldsymbol{\pi}} \} \sim \sum\limits_{k=1}^K{ \color{green}{\pi_k} \ \mathcal{GP}\Big(\mu_{\color{green}{k}}, \Psi_i^\color{green}{k} \Big)}, \ \perp \!\!\! \perp_i\]
Variational EM
E step: \[ \begin{align} \hat{q}_{\boldsymbol{\mu}}(\boldsymbol{\mu}) &= \color{green}{\prod\limits_{k = 1}^K} \mathcal{N}(\mu_\color{green}{k};\hat{m}_\color{green}{k}, \hat{\textbf{C}}_\color{green}{k}) , \hspace{2cm} \hat{q}_{\boldsymbol{Z}}(\boldsymbol{Z}) = \prod\limits_{i = 1}^M \mathcal{M}(Z_i;1, \color{green}{\boldsymbol{\tau}_i}) \end{align} \] M step:
\[ \begin{align*} \hat{\Theta} &= \underset{\Theta}{\arg\max} \sum\limits_{k = 1}^{K}\ \mathcal{N} \left( \hat{m}_k; \ m_k, \boldsymbol{C}_{\color{green}{\gamma_k}} \right) - \dfrac{1}{2} \textrm{tr}\left( \mathbf{\hat{C}}_k\boldsymbol{C}_{\color{green}{\gamma_k}}^{-1}\right) \\ & \hspace{1cm} + \sum\limits_{k = 1}^{K}\sum\limits_{i = 1}^{M}\tau_{ik}\ \mathcal{N} \left( \mathbf{y}_i; \ \hat{m}_k, \boldsymbol{\Psi}_{\color{brown}{\theta_i}, \color{brown}{\sigma_i^2}} \right) - \dfrac{1}{2} \textrm{tr}\left( \mathbf{\hat{C}}_k\boldsymbol{\Psi}_{\color{brown}{\theta_i}, \color{brown}{\sigma_i^2}}^{-1}\right) \\ & \hspace{1cm} + \sum\limits_{k = 1}^{K}\sum\limits_{i = 1}^{M}\tau_{ik}\log \color{green}{\pi_{k}} \end{align*} \]
Prediction
-
Multi-task posterior for each cluster:
\[ p(y_*(\mathbf{t}^{p}) \mid \color{green}{Z_{*k}} = 1, y_*(\mathbf{t}_{*}), \textbf{y}) = \mathcal{N} \Big( y_*(\mathbf{t}^{p}); \ \hat{\mu}_{*}^\color{green}{k}(\mathbf{t}^{p}) , \hat{\Gamma}_{pp}^\color{green}{k} \Big), \forall \color{green}{k}, \]
\(\hat{\mu}_{*}^\color{green}{k}(\mathbf{t}^{p}) = \hat{m}_\color{green}{k}(\mathbf{t}^{p}) + \Gamma^\color{green}{k}_{p*} {\Gamma^\color{green}{k}_{**}}^{-1} (y_*(\mathbf{t}_{*}) - \hat{m}_\color{green}{k} (\mathbf{t}_{*}))\)
\(\hat{\Gamma}_{pp}^\color{green}{k} = \Gamma_{pp}^\color{green}{k} - \Gamma_{p*}^\color{green}{k} {\Gamma^{\color{green}{k}}_{**}}^{-1} \Gamma^{\color{green}{k}}_{*p}\) -
Predictive multi-task GPs mixture:
\[p(y_*(\textbf{t}^p) \mid y_*(\textbf{t}_*), \textbf{y}) = \color{green}{\sum\limits_{k = 1}^{K} \tau_{*k}} \ \mathcal{N} \big( y_*(\mathbf{t}^{p}); \ \hat{\mu}_{*}^\color{green}{k}(\textbf{t}^p) , \hat{\Gamma}_{pp}^\color{green}{k}(\textbf{t}^p) \big).\]
An image is still worth many words

How about talent detection for swimmers?



Feel free to draw your own figures

Implemented as an R package MagmaClustR: https://github.com/ArthurLeroy/MagmaClustR
Not all problems are worth your time
A few examples of type 3 problems I have encountered over the years:
- Use longitudinal morphological or training information to provide better predictions.
- Make predictions on football (or other) games for online betting.
- Develop a (personalised) injury prevention tool using various training data.
- Identify optimal pacing strategies during a race.
-
Football. It’s whether purely random or obvious. Seriously, we should
just stop and move on.
Backup slides
\(\rightarrow\) All methods
scale linearly with the number of tasks.
\(\rightarrow\) Parallel computing
can be used to speed up training.
Overall, the computational
complexity is:
- Magma: \[ \mathcal{O}(M\times N_i^3 + N^3) \]
- MagmaClust: \[ \mathcal{O}(M\times N_i^3 + K \times N^3) \]
- Multi-Means Gaussian Processes: \[ \mathcal{O}(M \times P \times N_{ij}^3 + (M + P) \times N^3) \]
Appendix: MagmaClust, remaining clusters

Model selection performances of VBIC

Multi-Means Gaussian processes
Different sources of correlation might exist in the data (e.g. multiple genes and individuals)
\[y_{\color{blue}{i}\color{red}{j}} = \mu_{0} + f_\color{blue}{i} + g_\color{red}{j} + \epsilon_{\color{blue}{i}\color{red}{j}}\]
with:
- \(\mu_{0} \sim \mathcal{GP}(m_{0}, {C_{\gamma_{0}}}), \ f_{\color{blue}{i}} \sim \mathcal{GP}(0, \Sigma_{\theta_{\color{blue}{i}}}), \ \epsilon_{\color{blue}{i}\color{red}{j}} \sim \mathcal{GP}(0, \sigma_{\color{blue}{i}\color{red}{j}}^2), \ \perp \!\!\! \perp_i\)
- \(g_{\color{red}{j}} \sim \mathcal{GP}(0, \Sigma_{\theta_{\color{red}{j}}})\)
Key idea for training: define \(\color{blue}{M}+\color{red}{P} + 1\) different hyper-posterior distributions for \(\mu_0\) by conditioning over the adequate sub-sample of data.
\[p(\mu_0 \mid \{y_{\color{blue}{i}\color{red}{j}} \}_{\color{red}{j} = 1,\dots, \color{red}{P}}^{\color{blue}{i} = 1,\dots, \color{blue}{M}}) = \mathcal{N}\Big(\mu_{0}; \ \hat{m}_{0}, \hat{K}_0 \Big).\]
\[p(\mu_0 \mid \{y_{\color{blue}{i}\color{red}{j}} \}_{\color{blue}{i} = 1,\dots, \color{blue}{M}}) = \mathcal{N}\Big(\mu_{0}; \ \hat{m}_{\color{red}{j}}, \hat{K}_\color{red}{j} \Big), \ \forall \color{red}{j} \in 1, \dots, \color{red}{P}\]
\[p(\mu_0 \mid \{y_{\color{blue}{i}\color{red}{j}} \}_{\color{red}{j} = 1,\dots, \color{red}{P}}) = \mathcal{N}\Big(\mu_{0}; \ \hat{m}_{\color{blue}{i}}, \hat{K}_\color{blue}{i} \Big), \forall \color{blue}{i} \in 1, \dots, \color{blue}{M} \]
